The Computational Biology Research Center
The Computational Biology Research Center (CBRC) was founded in 2011 by Dr. Fatemeh Zare-Mirakabad , a faculty member of the Amirkabir University of Technology. The purpose of this center is to be a place for students and professors who are interested in computational topics of bioinformatics field to come together. This center Has played a significant role in promoting the scientific level of students and researchers by creating a relaxed and ambient environment. In this center, there are 14 bachelor students, 55 master students, 11 Ph.D. students and 4 researchers gathered to work and exchange up-to-date ideas in bioinformatics field.
Ongoing events

Weekly Seminars
Online - Weekly Seminars - Autumn & Winter 2021 - 2022
Time:
Wednesdays at 18:00 (GMT +3:30)
Topics:
- RNA
- Cancer
- Protein
- Drug Design
- Biological Network
- Reads Mapping
- Machine Learning in Bioinformatics
Research areas

Coronavirus disease
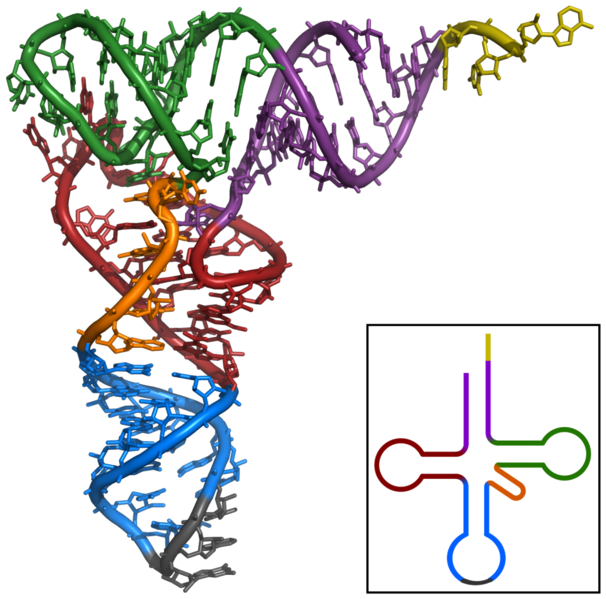
Structural bioinformatics
Drug
Sequence in bioinformatics

Biological network
In the news

Seminars
16 Mar, 2022

Arezou Pakseresht

Seminars
09 Mar, 2022

Mahdis Habibpour

Seminars
02 Mar, 2022

Zahra Seraj
Tutorials
Prediction mRNA subcellular localization
In post-transcriptional gene regulation, messenger RNA subcellular localization processes are critical. Trans-acting RNA-binding proteins interact with cis-regulatory regions called zipcodes to facilitate trafficking. While current sequencing-based technologies enable high-throughput identification of RNAs localized to specific subcellular compartments...

Protein Secondary Structure Prediction
Researchers have proposed over fifty years of approaches for predicting secondary structures of proteins in silico in the literature. Artificial neural network-based techniques, a subject of artificial intelligence and machine learning that is gaining increasing prominence in numerous application areas, make up a substantial part of this collection.

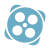
Autoencoder
In biology and bioinformatics, annotation of genomic data is a key difficulty. Existing databases of known gene functions are insufficient and prone to errors, and the bimolecular experiments required to update them are time-consuming and expensive. While computational methods cannot replace experimental verification, they can assist in two ways....
Softwares

ADRP-ML-NMF
Identifying and controlling adverse drug reactions is a complex problem in the pharmacological field. Despite the studies done in different laboratory stages, some adverse drug reactions are recognized after being released, such as Rosiglitazone. Due to such experiences, pharmacists are now more interested in using computational methods to predict adverse drug reactions.

DRP-VEM
Traditional drug discovery methods are costly and time-consuming. The use of the existing approved drugs for treating another disease is called drug repositioning, a common strategy to overcome traditional drug discovery issues.

TranDTA
TranDTA is the first method that applies transformers to extract feature of protein sequence and uses transformer representations in drug target binding affinity (DTBA) prediction.

CNDP_PPI_networks
The data set used in the paper contains the PPI networks of E. coli and S. cerevisiae which are extracted from the Database of Interacting Proteins (DIP). To label the proteins as essential/non-essential, the essential genes data of these species are collected from DEG database.