Warning: include(../header.html): Failed to open stream: No such file or directory in D:\Pleskvhosts\bioinformatics.aut.ac.ir\httpdocs\softwares\cebcp\index.php on line 47
Warning: include(): Failed opening '../header.html' for inclusion (include_path='.;.\includes;.\pear') in D:\Pleskvhosts\bioinformatics.aut.ac.ir\httpdocs\softwares\cebcp\index.php on line 47
Services
Towards Learning a Unified Latent Space for Bioactivity Detection
Motivation
Sometimes, drugs are withdrawn due to low bioactivity that is not observed during the experimental procedure. Thus, predicting bioactivity classes during the lead optimization reduces the failures risk and improves the enhancement of compound bioactivities. Recent studies demon-strate a relationship between the chemical structure of compounds and their bioactivity, i.e., struc-ture-activity relationship (SAR), without considering the complex relationship between drugs and bioactivity classes.
Results
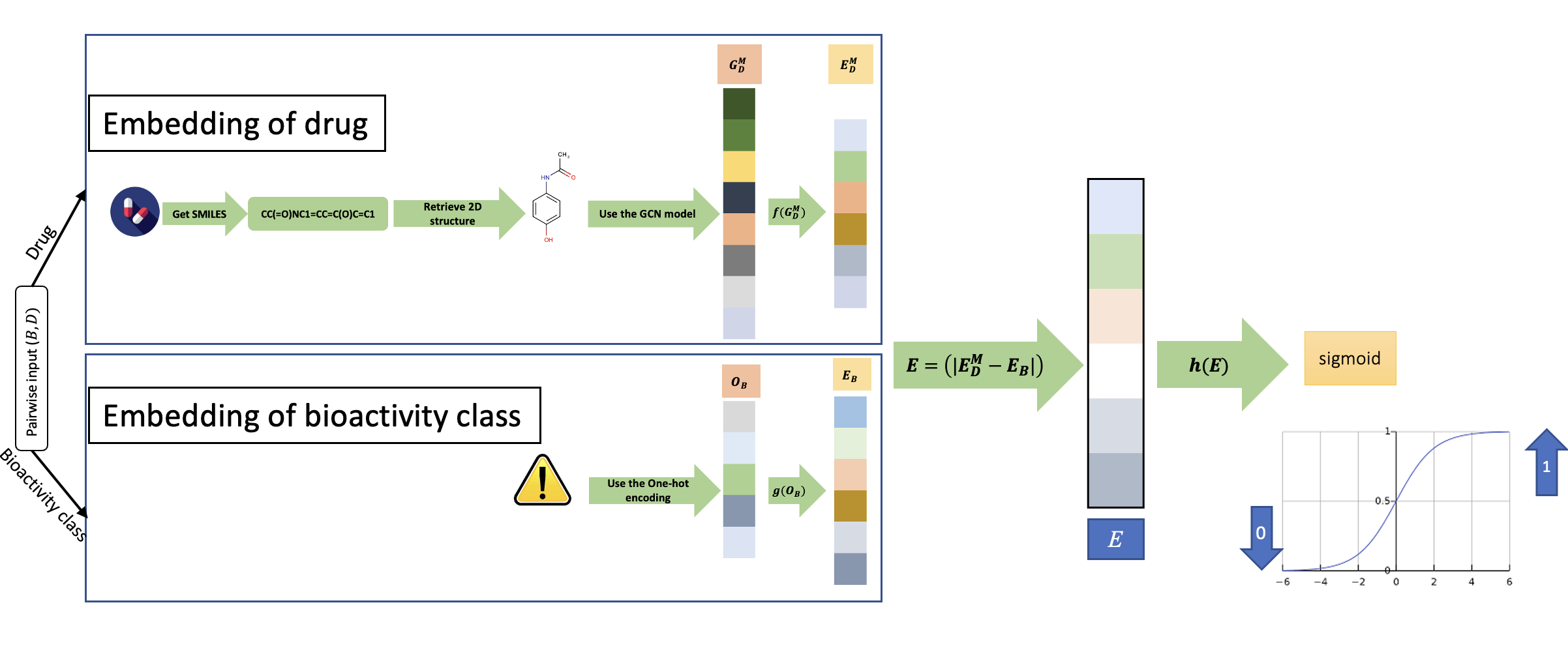
Results: We propose the Compositional Embedding for Bioactivity Class Prediction (CEBCP) method, which leverages compositional embedding and meta-learning to build a unified latent space for drugs and bioactivity classes. CEBCP utilizes the heterogenous siamese neural network as a meta-learning approach to understanding the similarity of drugs and bioactivities to tackle the limited data points. The CEBCP achieved the AUROC of 90.27%, 85.9%, and 83.25% by using association, drug, and bioactivity class strategies, respectively. The experimental results verified that the performance of the model could significantly outperform retrospective studies.
Availability
Availability: The data and code underlying this article are available in here. However, datasets were derived from sources in the public domain: https://sideeffects.embl.de .
Downloads
- Association_based_strategy.ipynb (266 KB).
- drug_based_strategy (176 KB).
- Bioactivity_class_based_strategy_(Meta_Learning).ipynb (527 KB).
- sider.csv (190 KB).
Warning: include(../footer.html): Failed to open stream: No such file or directory in D:\Pleskvhosts\bioinformatics.aut.ac.ir\httpdocs\softwares\cebcp\index.php on line 103
Warning: include(): Failed opening '../footer.html' for inclusion (include_path='.;.\includes;.\pear') in D:\Pleskvhosts\bioinformatics.aut.ac.ir\httpdocs\softwares\cebcp\index.php on line 103